Bioinformatics and insects… James Gillahan, GrowNextGen teacher leader and science instructor at Woodridge High School, recently attended a bioprospecting workshop aimed at training teachers to use authentic science research in the classroom. The educators engaged in hands-on laboratory practices and learned about navigating online genomics databases.
Gillahan’s participation was made possible by contributions from both GrowNextGen and the Six District Educational Compact. He shares about the experience:
The organizer of the teacher training was Seeds of Change, led by Beth Hunt and John Doleman, both of the Minneapolis, Minnesota area. Seeds of Change has developed several programs that involve a 10-day scientific research excursion to Costa Rica. Over the past few years, they have developed a vast citizen science project which includes a pure scientific research progression of labs that can be done in a high school lab setting.
This process starts with understanding the need for novel and effective antibiotics. The group has reason to believe that we may utilize the existing symbiotic and parasitic relationships between fungi, bacteria, and insects. Students can investigate what is known about these relationships and what is called the insect microbiome, which includes all bacteria that live in, on, and around the insect. After isolating any bacteria of interest from the insect microbiome, students complete a number of bioassays to elucidate any antibiotic properties from bacterial isolates.
If students find bacteria with antibiotic properties, the students can make tens of thousands of copies of its DNA right in our lab by taking advantage of what is called a polymerase chain reaction (PCR). Once there are enough copies of the DNA, the sample can be sent in for 16S genetic sequencing. The 16S region of DNA is found in all living organisms. It contains both highly conserved regions of DNA interspersed with unique regions that help distinguish one type of organism from another. Students will be able to upload their 16S sequence results to a database that compares it to all the other known genetic sequences (BLAST search).
From there, the student can construct a phylogenetic tree to show how closely their organism is related to others. A few organisms that seem particularly interesting or unique from all others might warrant being fully sequenced (whole genome sequencing). Results from the whole genome sequence might even merit further research by Seeds of Change partner researchers at the university level. This is what makes this program so exciting. Students might find something novel that could fight off a human pathogen that is otherwise resistant to the antibiotics currently available.
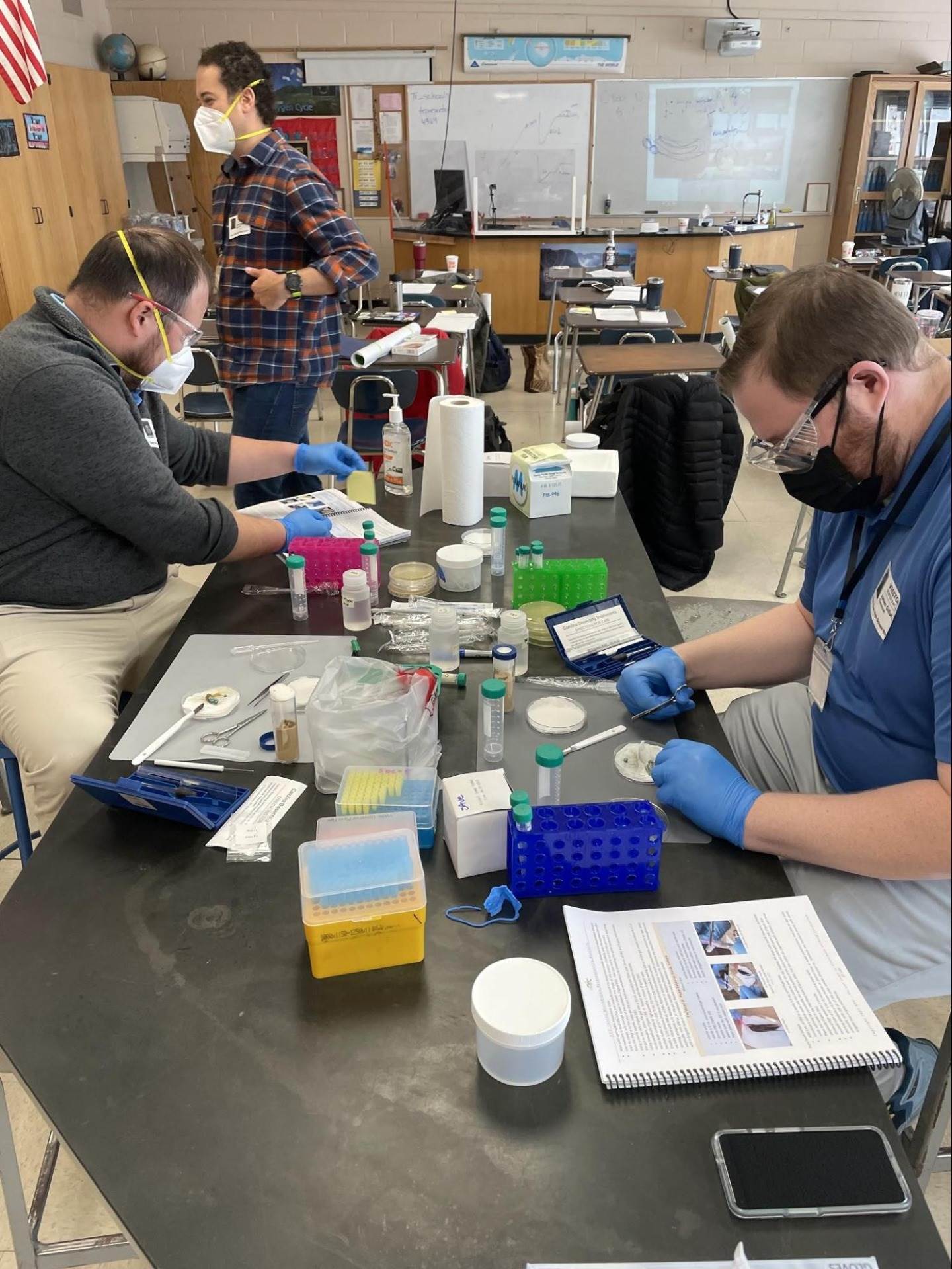

Students would also be able to use other bioinformatic tools to determine the presence and components of a cluster of genes responsible for producing an antibiotic or other biosynthetic molecule of interest. This sequence of autonomous experiments, research, and discovery is exciting, challenging, and relevant for students and for the teacher as well. This concept of exploiting an existing relationship between organisms to find a target drug or other compound of interest is central to bioprospecting. More fascinating still is using this technique to create a solution for the rise of antibiotic-resistant bacteria and dwindling antibiotics in clinical trials, as well as the classification of bacteria using bioinformatics—a perfect example of cutting edge biotechnology.
“I would love to start incorporating these labs into my level 2 capstone projects. I’d like to give my students the opportunity to start with a topic that affects virtually everyone, antibiotic resistance. Students would learn more about catalysts of resistance and what is currently being done, and then they will start researching, experimenting, and documenting a potential solution. After experiencing every part of the lab myself, seeing every step that has to be taken and the pitfalls and minor successes along the way, I am confident that a student that experiencing this much of the scientific process before graduating high school would be a fantastic start to anyone’s career.
“I look forward to continued collaboration with GrowNextGen, providing lesson materials, development, and/or demonstration of these labs to other Ohio science teachers through the Ohio Soybean Council.
“My thoughts on tying this together with an agricultural industry in Ohio: we could potentially use insects in and around farms where antibiotics are used. I was thinking these areas may already have antibiotic resistant bacteria around, so the bugs may have other antibiotic-producing bacteria that can out-compete the antibiotic-resistant bacteria to live (commensally, mutually, or pathogenetically) on the insect.
“It’s interesting to consider that insects are so much of a pest to both the soy and corn (plants) industry, we literally have developed transgenic plants that include a bacterial toxin that destroys the stomachs of insects (Bt corn/soy). It seems difficult to think how we could use insects and bacteria on them to help plants of interest, but I’d like to look for a way for us to link them together.”
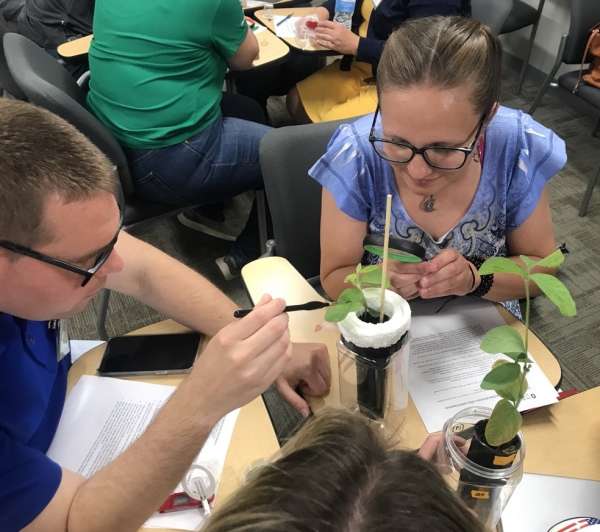
photos courtesy of Toms River Regional Schools




Share this